Structure Database (LMSD)
Common Name
PG(14:0/16:0(9Cp))
Systematic Name
1-tetradecanoyl-2-(9R, 11S-methylene-hexadecanoyl)-sn-glycero-3-phospho-(1'-sn-glycerol)
Synonyms
LM ID
LMGP04010991
Formula
Exact Mass
Calculate m/z
706.478488
Sum Composition
Status
Active
No other lipid differing only in stereochemistry/bond geometry found
3D model of PG(14:0/16:0(9Cp))
Please note: Where there are chiral atoms but the stereochemistry is undefined, the 3D model takes an arbitrary conformation
Main
Classification
Category
Main Class
Sub Class
String Representations
InChiKey (Click to copy)
GNWIBIDKLVNWES-OQIMXIDESA-N
InChi (Click to copy)
InChI=1S/C37H71O10P/c1-3-5-7-9-10-11-12-13-14-17-21-25-36(40)44-30-35(31-46-48(42,43)45-29-34(39)28-38)47-37(41)26-22-18-15-16-20-24-33-27-32(33)23-19-8-6-4-2/h32-35,38-39H,3-31H2,1-2H3,(H,42,43)/t32-,33+,34-,35+/m0/s1
SMILES (Click to copy)
[H][C@](O)(CO)COP(OC[C@]([H])(OC(CCCCCCC[C@@H]1C[C@@H]1CCCCCC)=O)COC(CCCCCCCCCCCCC)=O)(=O)O
References
Taxonomy Information
Curated from
NCBI taxonomy class
Reference
Escherichia coli
(#562)
Gammaproteobacteria
(#1236)
Lipid composition of membranes of Escherichia coli by liquid chromatography/tandem mass spectrometry using negative electrospray ionization.,
Rapid Commun Mass Spectrom, 2007
Rapid Commun Mass Spectrom, 2007
Pubmed ID:
17477452
Other Databases
PubChem CID
Calculated Physicochemical Properties
Heavy Atoms
48
Rings
1
Aromatic Rings
0
Rotatable Bonds
37
Van der Waals Molecular Volume
738.75
Topological Polar Surface Area
148.82
Hydrogen Bond Donors
3
Hydrogen Bond Acceptors
10
logP
10.85
Molar Refractivity
192.26
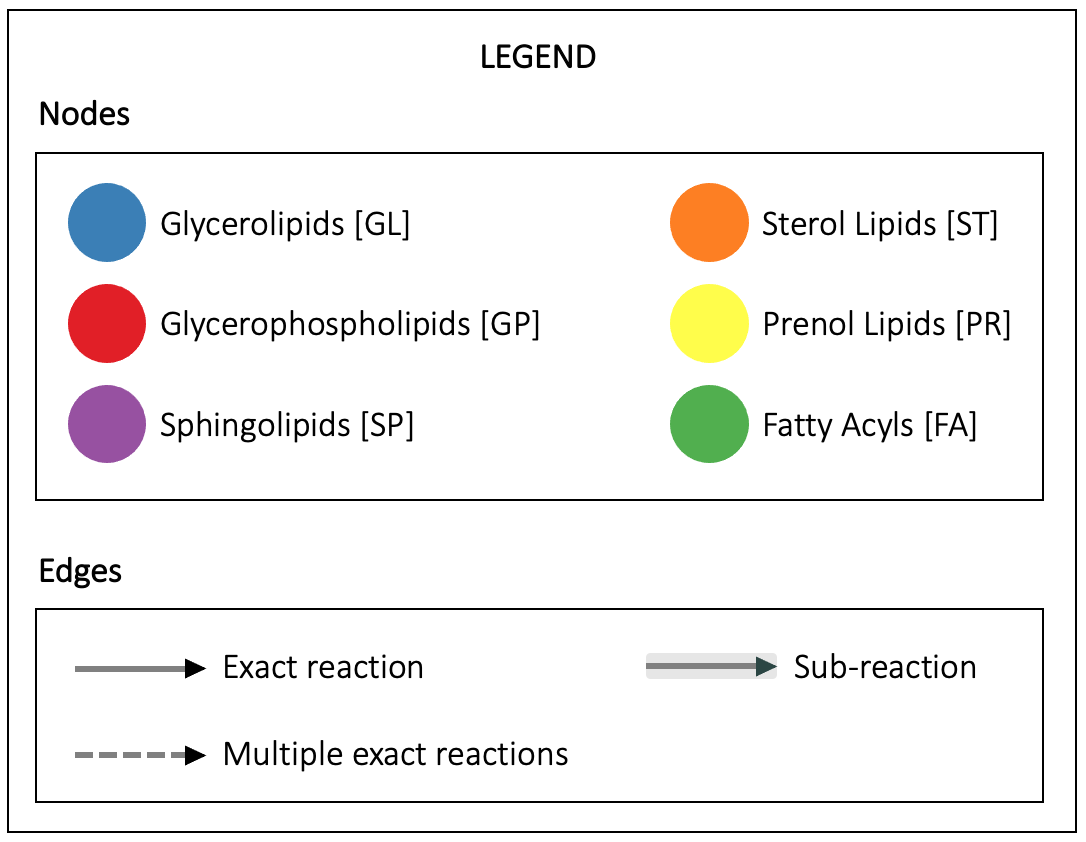
Reactions
Filter by species:
ⓘ
Reactions are shown if the E.C. number of the enzyme catalysing it is annotated in the UniProt database for a species belonging to the selected taxonomic class.
Click on an edge to display the reaction(s).
Admin
Created at
-
Updated at
25th Apr 2022
LIPID MAPS® abbreviations for glycerophospholipids (GP)
The LIPID MAPS® glycerophospholipid abbreviations (PC, PE, etc.) are used here to refer to species with one or two radyl side-chains where the structures of the side chains are indicated within parentheses in the 'Headgroup(sn1/sn2)' format (e.g. PC(16:0/18:1(9Z)). By default, R stereochemistry at the C-2 carbon of glycerol and attachment of the headgroup at the sn3 position is assumed. Also, acyl chains are assumed by default. The 'O-' prefix is used to indicate the presence of an alkyl ether substituent e.g. PC(O-16:0/18:1(9Z)), whereas the 'P-' prefix is used for the 1Z-alkenyl ether (Plasmalogen) substituent e.g. PC(P-16:0/18:1(9Z)).
For molecules with opposite (S) stereochemistry at C2 of the glycerol group and attachment of the headgroup at the sn1 position, the stereochemistry specification of [S] is appended to the abbreviation. The 'Headgroup(sn3/sn2)' abbreviation format is used.
For molecules with unknown stereochemistry at the C-2 carbon of the glycerol group, the stereochemistry specification of [U] is appended to the abbreviation and the structure is drawn with C-2 stereochemistry unspecified.
The LIPID MAPS® glycerophospholipid abbreviations (PC, PE, etc.) are used here to refer to species with one or two radyl side-chains where the structures of the side chains are indicated within parentheses in the 'Headgroup(sn1/sn2)' format (e.g. PC(16:0/18:1(9Z)). By default, R stereochemistry at the C-2 carbon of glycerol and attachment of the headgroup at the sn3 position is assumed. Also, acyl chains are assumed by default. The 'O-' prefix is used to indicate the presence of an alkyl ether substituent e.g. PC(O-16:0/18:1(9Z)), whereas the 'P-' prefix is used for the 1Z-alkenyl ether (Plasmalogen) substituent e.g. PC(P-16:0/18:1(9Z)).
For molecules with opposite (S) stereochemistry at C2 of the glycerol group and attachment of the headgroup at the sn1 position, the stereochemistry specification of [S] is appended to the abbreviation. The 'Headgroup(sn3/sn2)' abbreviation format is used.
For molecules with unknown stereochemistry at the C-2 carbon of the glycerol group, the stereochemistry specification of [U] is appended to the abbreviation and the structure is drawn with C-2 stereochemistry unspecified.